Open Access | Research Article
This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
Anti-tumor effect of pcDNA3.1(-)/rMETase-shUCA1 hyaluronic acid-modifed G5 polyamidoamine nanoparticles on gastric cancer
* Corresponding author: Chunying Xie
Mailing address: Department of Oncology, The Second Affiliated Hospital of Nanchang University, No. 1 Minde Road, Donghu District, Nanchang, China.
E-mail: 516467669@qq.com
Received: 14 May 2020 / Accepted: 09 June 2020
DOI: 10.31491/APT.2020.06.019
Abstract
Background: Recently, recombinant methioninase (rMETase) has been widely exploited as a chemotherapeutic
option during gastric cancer treatment. The present study utilized the pcDNA3.1(-)/rMETase-shUCA1 complex
and hyaluronic acid (HA)–modified fifth-generation polyamidoamine nanoparticles to evaluate the anti-tumor
function of the nanocarrier in Gastric cancer (GC) cells.
Methods: The characteristics of nanoparticles were analyzed using transmission electron microscopy. The GC
cell viability and invasion were tested using the CCK-8 assay and the cell invasion assay. The expressions of
CD44 phosphorylated mammalian target of rapamycin (p-mTOR), and c-caspase 3 were measured by Western
blot assay. Also, a nude mice xenograft model was established to assess the function of HA-polyamidoamineAu-METase-shUCA1 in GC tumor growth.
Results: The transfection of rMETase, alone and in combination with shUCA1 carried by HA-polyamidoamineAu, inhibited GC cell viability, invasion, and tumorsphere formation, and enhanced METase activity. Moreover,
HA-polyamidoamine-Au-METase-shUCA1 decreased p-mTOR expression and increased c-caspase 3, while this
effect could be reversed by mTOR activator MHY 1485. The CD44 positive cell number and the tumor volume in
nude mice, injected with HA-polyamidoamine-Au-METase-shUCA1were reduced.
Conclusions: The HA-polyamidoamine-Au-METase-shUCA1 nanoparticles restrained GC tumor growth, which
was partly via the mTOR pathway.
Keywords
Nanoparticles, gastric cancer, UCA1, rMETase
Introduction
Gastric cancer (GC) [1] remains one of the main reasons for cancer-related deaths [2] and presents with high
morbidity in China. During the past few decades, several
methods for the treatment of GC have made progress [3],
but the treatment effect of GC patients is still poor due to
the high metastasis of GC [4]. Clinically, recombinant methioninase (rMETase) has been widely used to treat GC [5], while resistance to rMETase causes therapeutic failure [6],
leading to the recurrence of GC and cancer-related death.
Therefore, investigating the molecular mechanism of GC
growth is urgently warranted.
Gene therapy can be an innovative therapeutic modality
for advanced GC [7]. In the evolution of gene therapies,
more and more genes have been proven that is a pivotal
role in GC treatment. Urothelial carcinoma-associated 1
(UCA1) is a long non-coding RNA (lncRNA) molecule
with three exons that encode a 1.4 kb and a 2.2 kb isoform
[8]. Several researchers have observed that UCA1 plays a
pivotal role in tumorigenesis through DNA rearrangements
or amplifications in several cancers, particularly of nonsmall cell lung cancer [9] and hepatocellular carcinoma
[10]. UCA1 was increased in GC cells, and a TGF-β1–induced UCA1 increase promoted GC cell invasion and migration [11]. Fang et al. indicated that UCA1 increased GC
multi-drug resistance by decreasing miR-27b [12]. Taken
together, UCA1 could be used as a potential target and a therapeutic strategy for GC. Nanoparticles have gradually
become a research hotspot due to their good biocompatibility, low toxicity in vivo, biodegradability, and easy
penetration of solid tumors [13, 14]. Moreover, appropriate chemical modifcation of nanoparticles could directly
and effectively kill tumor cells [15]. Polyamidoamine
dendrimers, which have low toxicity and non-immunogenicity characteristics, are used to combine molecules
to format gene delivery systems. As reported, hyaluronic
acid (HA) is a receptor for CD44, which is mainly characterized by good biocompatibility and water solubility [16]
and is related to tumor invasion and metastasis [17]. Thus,
HA has been widely used as a tumor cell modifier. The
half-life of drugs in the blood was also elevated through
nanocarriers and reduced HA modifcation. In the current
study, we created HA-modifed ffth-generation polyamidoamine nanoparticles loaded with UCA1 small hairpin
RNA (shRNA) and Au-METase (HA-polyamidoamineAu-METase-shUCA1), and we explored the functional
role of HA-polyamideamine-Au-METase-shUCA1 in GC
cell viability and invasion, which might provide a foundation for research into the treatment of GC.
Materials and Methods
Nanoparticle preparation
HA (Lingbao Biological Technology Co., Ltd., Beijing, China) and G5 polyamidoamine (Sigma, USA) were lysed in sodium borohydride (NaBH4) buffer with a concentration of 0.1 M and a pH of 8.5. Then, the mixture was catalyzed under NaBH3CN at 40°C for 3 days to make HApolyamidoamine polymer with 5% grafting density. The reaction solution was then dialyzed in a dialysis container. Then, the previously prepared HA-polyamidoamine and chloroauric acid (HAuCl4) were mixed at a mole ratio of 25:1, and NaBH4 was added to this compound. Gold (Au) nanoparticles were packed into the interior cavities of the HA-polyamide amide to result in HA-polyamidoamineAU. The HA-polyamidoamine-Au complex was lysed via phosphate-buffered saline (PBS), and then we appended a plasmid-containing METase (pcDNA-METase; Shionogi & Co., Ltd, Japan) to this solution, and then cultivated it for about 30min. Agarose gel electrophoresis (AGE) was performed to estimate HA-polyamidoamine-Au-METase effciency.
Morphological observation of nanoparticle characterization
We used transmission electron microscopy (TEM) to observe nanoparticle morphology. In brief, we determined the average particle size and size distribution using a particle size distribution tester.
Cell culture
We purchased Human GC cell line NCI-N87 from American Type Culture Collection. NCI-N87 cells were placed in Dulbecco’s Modifed Eagle Medium (DMEM; Sigma, USA) with the addition of 10% (v/v) heat-inactivated fetal bovine serum (FBS, Sigma, USA), 2 mM glutamine, 100units/mL penicillin, and 100 μg/mL streptomycin at a humidifed surrounding of 37°C, 5% carbon dioxide (CO2).
Isolation of NCI-N87 cancer stem cells
We collected NCI-N87 cells and then cultured them in serum-free high glucose DMEM (20 ng/mL epidermal growth factor, 10 ng/mL basic fibroblast growth factor). When the NCI-N87 cells were cultured in a spherical shape, we used a magnetically-activated cell sorting (MACS) system to sort CD44 positive cells. Next, we gathered spheroids and gained NCI-N87 cancer stem cells (CSCs) using the magnetic beads of the MACS.
Cell transfection
HA-polyamidoamine-Au, HA-polyamidoamine-AuMETase, HA-polyamidoamine-Au-shUCA1, and HApolyamidoamine-Au-METase-shUCA1 nanoparticles were transfected into NCI-N87 or NCI-N87 CSCs using Lipofectamine 2000 (Invitrogen, San Diego, CA, USA).
Cell viability assay
Accutase® cell dissociation buffer (Life Technologies Corporation) was used to dissociate NCI-N87 or NCIN87 CSCs, and then we inoculated these cells (1.0×103 cells) into 96-well microplates. Based on the manufacturer’s instructions, a volume of 10 μL CCK-8 was placed in each well. Next, the survival and viability of the above differently treated cells were recorded by determining the absorbance at 450 nm with a Spectra Max M5 microplate reader.
Cell invasion experiment
We placed NCI-N87 or NCI-N87 CSCs in the upper chamber (pore size of 8 μm, Corning) with 40 μL Matrigel (2 mg/mL), and then we placed 600 μL DMEM with 20% FBS in the lower chamber. After incubating for 1 day, we stained the membrane with 2% crystal violet solution for about 10 min. Ultimately, we used a microscope to count in 5 random fields. All determinations were performed three independent times.
METase activity assay
Briefly, GC cells transfected with different vectors were collected and sonicated for 1 min and centrifuged. Subsequently, the activity of METase in the supernatant after centrifugation was measured by quantifying the a-ketobutyrate produced by 10 mM methionine with 3-methyl- 2-benzothiazoline hydrazone. Ultimately, we used a Hitachi U-2000 spectrophotometer (Beijing, China) to quantify the product at an optical density (OD) of 335 nm.
RNA isolation and qRT-PCR assay
Based on the manufacturer’s standard procedure, we separated total RNA from cells, and then we reversetranscribed complementary DNA (cDNA) via a reversetranscription kit. Next, we performed a real-time polymerase chain reaction (PCR) experiment upon a 7500 Fast Dx Real-Time PCR Instrument (Applied Biosystems) (Thermo Fisher Scientific, USA) instrument by adding SYBR Green PCR Master Mix (Thermo Fisher Scientifc, USA). The 2-∆∆CT method was performed to measure UCA1 expression.
Western blot analysis
Radioimmunoprecipitation assay (RIPA) buffer with protease inhibitors (Beyotime Institute of Biotechnology, Shanghai, China) was used to separate the total proteins. Next, we subjected these samples to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) and then transferred them into polyvinylidene fluoride (PVDF) membranes, and the membranes were then incubated with each primary antibody from Abcam (UK), including CD44, c-caspase 3, phosphorylated mammalian target of rapamycin (p-mTOR), total (t)-mTOR, and β-actin overnight in a 4 °C environment. Next, these membranes were incubated with the secondary antibody from Abcam for 2 h. Next, we visualized the immunoblots via an enhanced chemiluminescence (ECL) kit (Santa Cruz Biotechnology, USA).
Nude mice xenograft models
A total of 40 7-week-old BALB/c nude mice (female) were purchased from the Experimental Animal Center of Shanghai Cancer Institute. We performed these experiments based on the national regulations, and these experiments were approved by the ethical committee of The Second Affiliated Hospital of Nanchang University. We induced gastric cancer by inoculating mice in the flank with NCI-N87 CSCs (1×107 cells). HA-polyamidoamineAu, HA-polyamidoamine-Au-METase, HA-polyamidoamine-Au-shUCA1, and HA-polyamidoamine-AuMETase-shUCA1 were injected into the caudal vein of the mice on day 2, day 5, day 10, and day 15 (n = 10 per group). Ultimately, we killed the mice with the tumor on the 30th day and the equation: (volume [mm3] = length × width 2/2) was used to quantify the tumor volume. The percentage of CD44 positive cells in the tissue samples was examined using immunohistochemistry. In brief, we counted the number of CD44 positive cells in 5 randomly chosen high power felds of each group, and the ratio of other groups to the HA-polyamidoamine-Au group was calculated. The negative control was a section that used PBS instead of the primary antibody.
Statistical analysis
Data were exhibited as mean ± standard error mean, and all data were appraised with Student’s t-test or analysis of variance (ANOVA). SPSS 18.0 software was used to appraise these statistical analyses. A p-value of less than 0.05 indicates statistical signifcance.
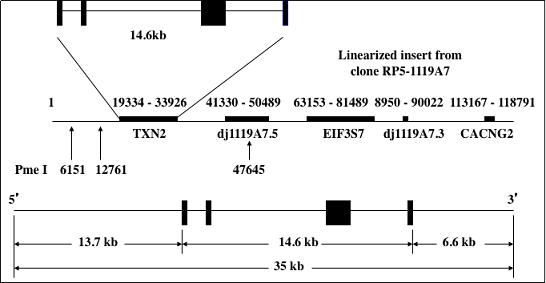
Figure 1. DNA construct for Tg(TXN2)+/0 mice. The human thioredoxin 2 gene (TXN2) with 13.7 kb and 6.6 kb of the 5’- and 3’-flanking sequences was used to generate Tg(TXN2)+/0 mice by pronuclear microinjection of zygotes from the mating of (C57BL/6J X SJL/J)F1 females with (C57BL/6J X SJL/J)F1 males.
Results
The characteristics of the nanoparticles
TEM was performed to observe the morphological features of HA-polyamidoamine-Au. The results expounded that the particle size distribution of Au nanoparticles was about 50 nm. The HA-polyamidoamine-Au carrier presented irregular spheres and good dispersion (Figure 1A), and HA-polyamidoamine-Au-METase-shUCA1 zeta potential was around 10.7 mV (Figure 1B). The particle size of HA-polyamidoamine-Au-METase-shUCA1 was about 206 nm (Figure 1C and 1D). Also, compared with the HApolyamidoamine-Au group, the particle size (nM) was increased in the HA-polyamidoamine-Au-METase, HApolyamidoamine-Au-shUCA1, and HA-polyamidoamineAu-METase + shUCA1 groups, while zeta potential (mV) was reduced (Figure 1E).
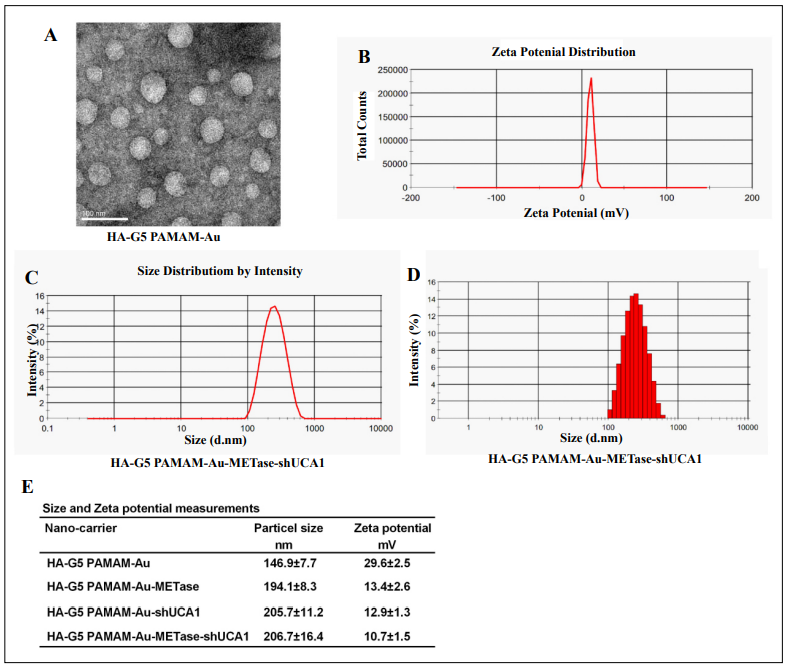
Figure 1. The characteristics of nanoparticles. (A) Transmission electron microscopy image of HA-polyamidoamine-Au. (B) Zeta potential distribution of HA-polyamidoamine-Au-METase-shUCA1. (C, D) Size distribution based on intensity of HA-polyamidoamine-Au-METase-shUCA1. (E) Statistical results of particle size (nM) and zeta potential (mV) of nanocarriers. G5 PAMAM: fifth- generation polyamidoamine; METase: methioninase; HA: hyaluronic acid.
HA-polyamidoamine-Au-METase-shUCA1 restrained METase activity and UCA1 expression in NCI-N87 and NCI-N87 CSCs
NCI-N87 cells and NCI-N87 CSCs were collected, and then we measured the CD44 expression and cell viability, respectively. The CD44 protein level was higher in NCIN87 CSCs (Figure 2A). CCK-8 assay indicated that when the concentration is less than 200 μg/mL, the carrier (HApolyamidoamine-Au) showed no prominent influence on cell viability. Once the concentration reached or exceeded 200 μg/mL, the cell viability was restrained not only in NCI-N87 but also in NCI-N87 CSCs (Figure 2B). After the transfection of HA-polyamidoamine-Au-METase or HA-polyamidoamine-Au-METase-shUCA1 for 3 days in NCI-N87 or NCI-N87 CSCs, the METase activity in the supernatant fluid was increased (Figure 2C), while the level of free MET was decreased (Figure 2D). Moreover, the relative UCA1 level was signifcantly reduced in NCIN87 and NCI-N87 CSCs with the transfection of HApolyamidoamine-Au-shUCA1 and HA-polyamidoamineAu-METase-shUCA1 (Figure 2E).
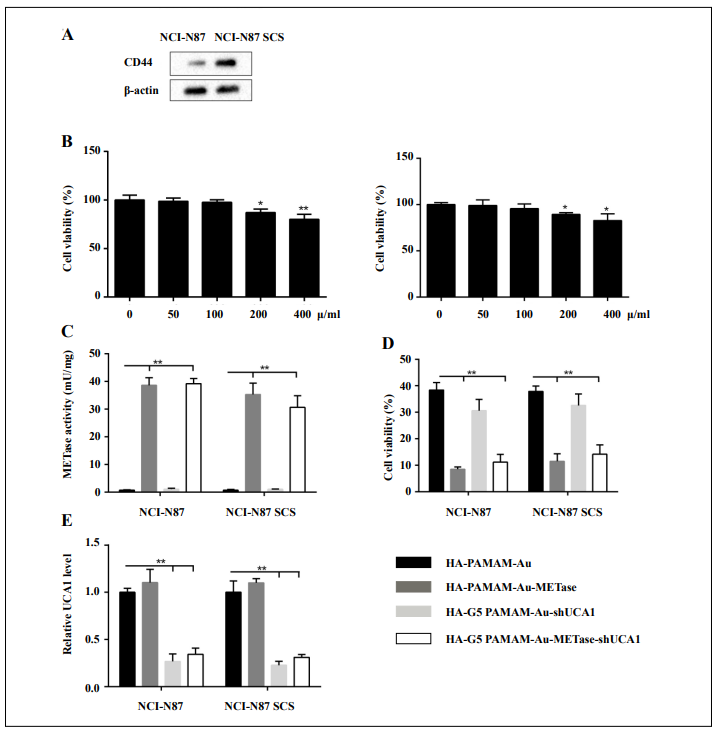
Figure 2. HA-polyamidoamine-Au-METase-shUCA1–inhibited METase activity and UCA1 expression in NCI-N87 and NCI-N87 cancer stem cells (CSCs). (A) The expression of CD44 in NCI-N87 and NCI-N87 CSCs was detected by Western blot analysis. β-actin acted as the internal control. (B) The effect of HA-polyamidoamine-Au on cell viability was detected by CCK- 8 assays. (C) METase activity transfected with different nanoparticles was assessed. (D) The effect of nanocarriers on free MET expression. (E) The relative UCA1 level in cells was detected by qRT-PCR. GAPDH was used as the internal control. *P < .05, **P < .01: comparison and analysis were conducted between groups. PAMAM: polyamidoamine; METase: methioninase; HA: hyaluronic acid.
HA-polyamidoamine-Au-METase-shUCA1–inhibited NCI-N87 cell viability and invasion
To expound the influence of nanoparticles on NCI-N87 cells, these cells were stochastically divided into six groups: HA-polyamidoamine-Au (which acted as a control), HA-polyamidoamine-Au-METase, HA-polyamidoamine-Au-shUCA1, HA-polyamidoamine-Au-METaseshUCA1, HA-polyamidoamine-Au-METase-shUCA1 + MHY 1485 (mTOR activator), and HA-polyamidoamineAu-METase-shUCA1 + Rapamycin (mTOR inhibitor). Results presented that the best suppression of NCI-N87 cell invasion can be achieved in the HA-polyamidoamineAu-METase-shUCA1 group. However, the number of invasive cells was markedly increased in the HA-polyamidoamine-Au-METase-shUCA1 + MHY 1485 group, as compared with the HA-polyamidoamine-Au-METaseshUCA1 group, and this effect could be reversed by Rapamycin supplementation (Figure 3A and 3B). Meanwhile, the outcome of cell viability presented a similar trend as cell invasion (Figure 3C). Western blot analysis revealed that three vectors (HA-polyamidoamine-AuMETase, HA-polyamidoamine-Au-shUCA1, and HApolyamidoamine-Au-METase-shUCA1) inhibited the p-mTOR expression and promoted c-caspase 3 levels; nevertheless, this effect was alleviated in cells transfected with HA-polyamidoamine-Au-METase-shUCA1 and MHY 1485, and the opposite results were presented in the HA-polyamidoamine-Au-METase-shUCA1 + Rapamycin group (Figure 3D).
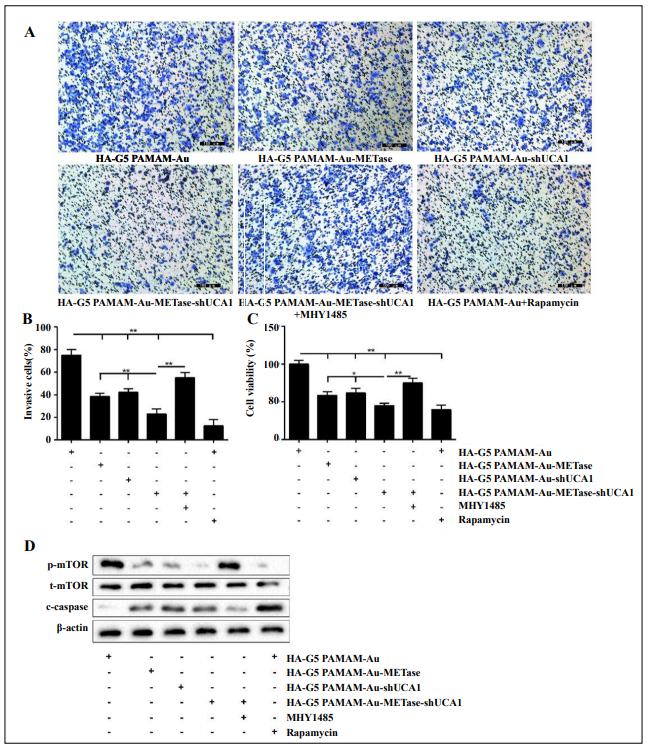
Figure 3. HA-polyamidoamine-Au-METase-shUCA1–inhibited NCI-N87 cell viability and invasion. (A) Representative images and (B) accompanying statistical plots were presented about cell invasion. (C) HA-polyamidoamine-Au-METase-shUCA1 suppressed NCI-N87 cell viability through CCK-8 assay. (D) The effect of different nanocarriers on the expression of p-mTOR and c-caspase 3 was detected through Western blot analysis. β-actin was used as the internal control. *P < .05, **P < .01: comparison and analysis were conducted between groups. G5 PAMAM: ffthgeneration polyamidoamine; METase: methioninase; HA: hyaluronic acid.
HA-polyamidoamine-Au-METase-shUCA1–inhibited NCI-N87 CSC viability and invasion
Similarly, to expound the influence of nanoparticles on NCI-N87 CSCs, the groups were the same as specified above. Three vectors (HA-polyamidoamine-Au combined with either METase or shUCA1or METase-shUCA1) suppressed cell invasion, and HA-polyamidoamine-AuMETase-shUCA1 showed the strongest effect. The same effect on cell invasion inhibition was also presented in the HA-polyamidoamine-Au-METase-shUCA1 + Rapamycin group, while this effect was mitigated by adding mTOR activator MHY 1485 (Figure 4A). A histogram presenting the data is shown in Figur 4B. Analogously, the results of cell viability were similar to those of cell invasion ability (Figure 4C). Besides, the level of p-mTOR was suppressed and c-caspase 3 was increased in the HApolyamidoamine-Au-METase, HA-polyamidoamine-AushUCA1, and HA-polyamidoamine-Au-METase-shUCA1 groups in comparison to the control group, while this situation was reversed in the HA-polyamidoamine-AuMETase-shUCA1 + MHY 1485 group as compared with the HA-polyamidoamine-Au-METase-shUCA1 group,and compared with the HA-polyamidoamine-Au-METaseshUCA1 + MHY 1485 group, the level of p-mTOR was suppressed and c-caspase 3 was increased in the HA-polyamidoamine-Au-METase-shUCA1 + Rapamycin group (Figure 4D).
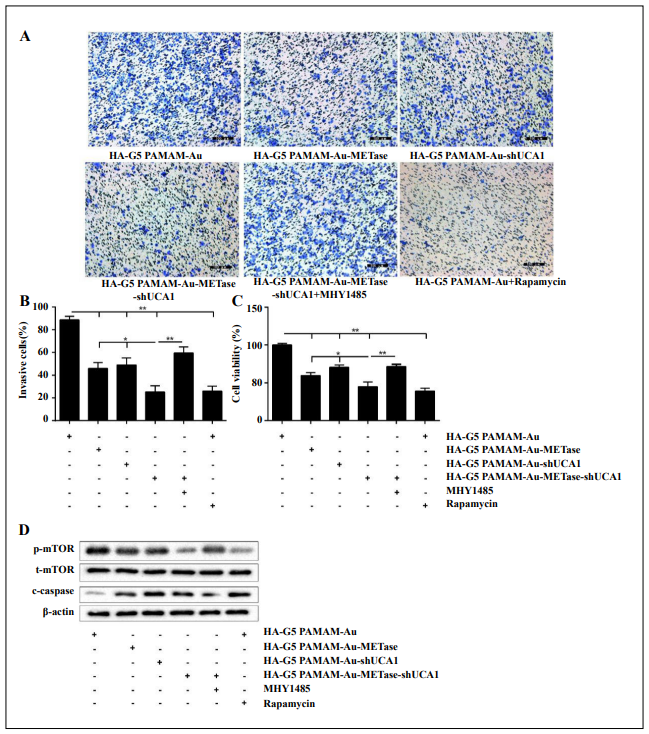
Figure 4. HA-polyamidoamine-Au-METase-shUCA1–inhibited NCI-N87 cancer stem cell (CSC) viability and invasion. Quantifcation of the invasion of NCI-N87 CSCs in groups according to METase and UCA1 gene expression. (A) Representative images and (B) accompanying statistical plots were demonstrated. (C) A CCK-8 assay was used to determine the effect of various nanoparticles on NCI-N87 CSC viability. (D) Protein levels of p-mTOR and caspase-3 after transfection were determined by Western blot analysis. β-actin was used as the internal control. *P < .05, **P < .01: comparison and analysis were conducted between groups. G5 PAMAM: ffth-generation polyamidoamine; METase: methioninase; HA: hyaluronic acid.
HA-polyamidoamine-Au-METase-shUCA1–inhibited tumor growth in vivo
Nanoparticles (HA-G5 polyamidoamine-Au, HA polyamidoamine-Au-METase, HA-polyamidoamine-Au-shUCA1, and HA-polyamidoamine-Au-METase-shUCA1) were injected into the caudal vein of the NCI-N87 CSC xenograft mouse model. As demonstrated in Figure 5A, the tumor volume of mice transfected with HA-polyamidoamineAu-METase, HA-polyamidoamine-Au-shUCA1, and HApolyamidoamine-Au-METase-shUCA1 was reduced. Furthermore, the immunohistochemistry results revealed that compared with the control group, the CD44 positive cell number in mice transfected with HA-polyamidoamineAu-METase, HA-polyamidoamine-Au-shUCA1, and HApolyamidoamine-Au-METase-shUCA1 was decreased (Figure 5B). Also, p-mTOR expression was accordingly reduced and c-caspase 3 expression was increased in the HA-polyamidoamine-Au-METase, HA-polyamidoamineAu-shUCA1, and HA-polyamidoamine-Au-METaseshUCA1 groups (Figure 5C).
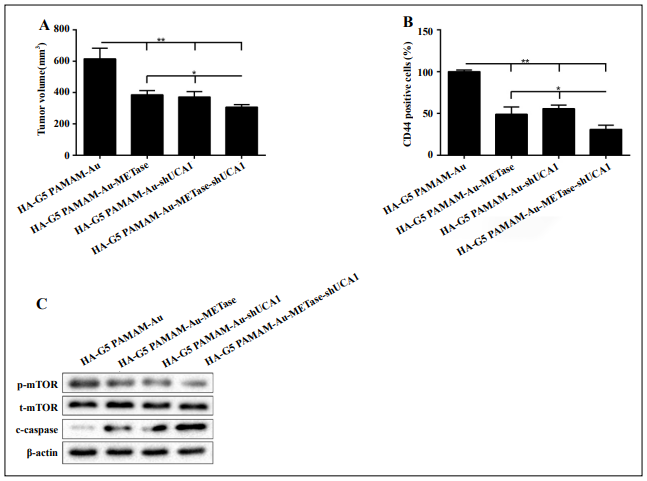
Figure 5. HA-polyamidoamine-Au-METase-shUCA1–inhibited tumor growth in mice induced with gastric cancer. Subcutaneous gastric cancer was induced via inoculation in the flank of 1 × 107 mouse NCI-N87 cancer stem cells (CSCs). (A) The tumor volume of caudal vein implantation models of NCI-N87 CSCs is shown. (B) The percentage of CD44 positive cells was determined using immunohistochemistry. (C) The expression of mTOR, p-mTOR, and c-caspase-3 was determined using Western blot analysis. β-actin was used as an endogenous control. *P < .05, **P < .01: comparison and analysis were conducted between groups. G5 PAMAM: ffth-generation polyamidoamine; METase: methioninase; HA: hyaluronic acid.
Discussion
The mortality associated with GC is a major problem
worldwide [18]. However, conventional treatment strategies for GC are not yet adequate. The purpose of this research was to evaluate the effects of HA-polyamidoamine
nanoparticles loaded with METase-shUCA1 on the growth
and anti-tumor function for GC in NCI-N87 and NCI-N87
CSCs.
GC is a major cause of cancer death in Asia [19]. Clinically, routine procedures, such as surgery and chemotherapy,
are used to relieve GC. Unfortunately, the therapeutic effects of those methods are not optimistic and chemotherapeutic agents also have many side effects, including drug
resistance [20]. Moreover, previous studies showed that
the resistance of chemotherapy was bound up with CSCs
with evidence that drug-resistant cancers were accompanied by increased CSC transcription factor expressions
[21]. As reported, the overexpressed CD44 in CSCs leads
to higher tumorigenicity of the cells and can lead to the
development of heterogeneous cancers [22]. In our study,
homologous gastric CSCs showed an increased CD44
level and were accompanied by higher migration and the
capability of tumorsphere formation.
In the treatment of diverse cancers, rMETase is widely
used as a therapeutic option because of its strong MET depletion abilities [23]. Tan et al. found that rMETase, alone,
and combined with cisplatin had a prominent anti-tumor
effect in colon cancer [24]. Besides, previous studies have
shown that rMETase-loaded stealth polylactic-co-glycolic
acid (PLGA)/liposomes modifed with anti-CAGE singlechain fragment variable (scFV) were effective for treating GC [25]. Here, we discovered that the overexpression of
rMETase boosted METase activity, restrained free MET
expression, and suppressed GC cell viability and invasion
that resulted in the increased expression of the apoptosisrelated c-caspase 3 protein.
LncRNAs are evolutionarily conserved non-protein-coding RNAs with a length of more than 200 nucleotides [26].
UCA1 is a lncRNA that bounds up with the concoction of
drug resistance in various tumors. For example, knockout
UCA1 increased how susceptive breast cancer cells were
to tamoxifen susceptibleness of breast cancer cells via
the restraint of the Wnt/β-catenin axis [27]. UCA1 also
increases acquired resistance to Epidermal Growth Factor Receptor (EGFR)- tyrosine kinase inhibitors (TKIs) in
EGFR-mutant non-small cell lung cancer by boosting the
AKT/mTOR axis [28]. The above findings suggest that
UCA1 might be a momentous lncRNA modulator of drug
resistance. However, the potential molecular mechanisms
remain largely unknown. Here, we compared UCA1 expression between GC NCI-N87 and NCI-N87 CSCs and
expounded that silencing UCA1 in NCI-N87 and NCIN87 CSCs significantly lowered UCA1 expression. Our
data showed that the knockdown of UCA1 in both NCIN87 and NCI-N87 CSCs signifcantly decreased the cell
viability and invasion, and increased the expression of ccaspase 3.
Previous studies reported that a UCA1 increase can lead
to the activation of the mTOR axis in multiple types of
cancer cells [29]. For example, UCA1 conferred breast
cancer cell tamoxifen resistance partly through the mTOR
signaling axis [8]. In the evolution of non-small cell lung
cancer, UCA1 may induce non-T790M resistance of
EGFR-TKIs by activating the mTOR axis and epithelialmesenchymal transition [1]. Therefore, we hypothesized
that UCA1 might regulate GC progression via the mTOR
axis. The experimental data in this study expounded that
UCA1 small interfering RNA significantly decreased pmTOR in NCI-N87 and NCI-N87 CSCs, and restrained
cell viability and invasion. The above results indicate
that UCAI may be able to activate the mTOR signaling
pathway in GC cells. Our functional study showed that
MHY1485, an mTOR activator, significantly impacted
the above effect of UCA1 on GC cells, suggesting that the
UCA1-modulated GC process occurs at least partly via
the mTOR signaling pathway.
With the rapid development of nanotechnology in recent
years, more and more nanoparticle and microparticle
drug delivery systems generated from biodegradable
polymers have been used for the treatment of several cancers [30, 31]. The advantage of biodegradable synthetic
microparticles is mainly to regulate the drug release rate
by altering the polymer structure and biodegradation rate
to achieve the desired therapeutic concentrations [32].
Moreover, polyamidoamine dendrimers have displayed
great potential during the process of delivering drugs and
genes, owing to their high transfection effciency and low
toxicity [33]. HA-polyamidoamine was benefcial to avoid
macrophage phagocytosis through the hydrophilic layer
of nanoparticles [34]. In the current study, transfection of
rMETase, alone and in combination with UCA1 shRNA,
carried by HA-polyamidoamine restrained GC cell proliferation and invasion and boosted c-caspase 3 expressions.
In conclusion, HA-polyamidoamine-Au-METase-shUCA1
restrained GC tumor growth through the mTOR pathway,
suggesting that nanoparticle-mediated gene therapy can
provide a safe and effective therapeutic option for GC, although more evidence is still needed.
Declaration
Conflict of Interest
The authors declare that they have no conflict of interest.
References
1. Cheng N, Cai W, Ren S, et al. Long non-coding RNA UCA1 induces non-T790M acquired resistance to EGFR-TKIs by activating the AKT/mTOR pathway in EGFR-mutant nonsmall cell lung cancer. Oncotarget, 2015, 6(27): 23582.
2. Shen Y H, Xie Z B, Yue A M, et al. Expression level of microRNA-195 in the serum of patients with gastric cancer and its relationship with the clinicopathological staging of the cancer. Eur Rev Med Pharmacol Sci, 2016, 20(7): 1283-1287.
3. Peng W, Si S, Zhang Q, et al. Long non-coding RNA MEG3 functions as a competing endogenous RNA to regulate gastric cancer progression. Journal of experimental & clinical cancer research, 2015, 34(1): 79.
4. Shirong C, Jianhui C, Chuangqi C, et al. Survival of proper hepatic artery lymph node metastasis in patients with gastric cancer: implications for D2 lymphadenectomy. PloS one, 2015, 10(3).
5. Xin L, Cao J, Cheng H, et al. Stealth cationic liposomes modified with anti-CAGE single-chain fragment variable deliver recombinant methioninase for gastric carcinoma therapy. Journal of nanoscience and nanotechnology, 2013, 13(1): 178-183.
6. Yano S, Li S, Han Q, et al. Selective methioninase-induced trap of cancer cells in S/G2 phase visualized by FUCCI imaging confers chemosensitivity. Oncotarget, 2014, 5(18): 8729.
7. Zhu X, Su D, Xuan S, et al. Gene therapy of gastric cancer using LIGHT-secreting human umbilical cord bloodderived mesenchymal stem cells. Gastric Cancer, 2013, 16(2): 155-166.
8. Wu C, Luo J. Long non-coding RNA (lncRNA) urothelial carcinoma-associated 1 (UCA1) enhances tamoxifen resistance in breast cancer cells via inhibiting mTOR signaling pathway. Medical science monitor: international medical journal of experimental and clinical research, 2016, 22: 3860.
9. Nie W, Ge H, Yang X, et al. LncRNA-UCA1 exerts oncogenic functions in non-small cell lung cancer by targeting miR-193a-3p. Cancer letters, 2016, 371(1): 99-106.
10. Xiao J N, Yan T H, Yu R M, et al. Long non-coding RNA UCA1 regulates the expression of Snail2 by miR-203 to promote hepatocellular carcinoma progression. Journal of cancer research and clinical oncology, 2017, 143(6): 981-990.
11. Zuo Z K, Gong Y, Chen X H, et al. TGFβ1-induced lncRNA UCA1 upregulation promotes gastric cancer invasion and migration. DNA and cell biology, 2017, 36(2): 159- 167.
12. Fang Q, Chen X Y, Zhi X T. Long non-coding RNA (LncRNA) urothelial carcinoma associated 1 (UCA1) increases multi-drug resistance of gastric cancer via downregulating miR-27b. Medical science monitor: international medical journal of experimental and clinical research, 2016, 22: 3506.
13. Wang X, Teng Z, Wang H, et al. Increasing the cytotoxicity of doxorubicin in breast cancer MCF-7 cells with multidrug resistance using a mesoporous silica nanoparticle drug delivery system. International journal of clinical and experimental pathology, 2014, 7(4): 1337.
14. Abbasi S, Paul A, Shao W, et al. Cationic albumin nanoparticles for enhanced drug delivery to treat breast cancer: preparation and in vitro assessment. Journal of drug delivery, 2012, 2012.
15. You J, Zhang R, Xiong C, et al. Effective photothermal chemotherapy using doxorubicin-loaded gold nanospheres that target EphB4 receptors in tumors. Cancer research, 2012, 72(18): 4777-4786.
16. Ganesh S, Iyer A K, Gattacceca F, et al. In vivo biodistribution of siRNA and cisplatin administered using CD44- targeted hyaluronic acid nanoparticles. Journal of controlled release, 2013, 172(3): 699-706.
17. Wang Z, Shi Q, Wang Z, et al. Clinicopathologic correlation of cancer stem cell markers CD44, CD24, VEGF and HIF-1α in ductal carcinoma in situ and invasive ductal carcinoma of breast: an immunohistochemistry-based pilot study. Pathology-Research and Practice, 2011, 207(8): 505-513.
18. Peng Y, Zhang X, Ma Q, et al. MiRNA-194 activates the Wnt/β-catenin signaling pathway in gastric cancer by targeting the negative Wnt regulator, SUFU. Cancer letters, 2017, 385: 117-127.
19. Kim Y H, Liang H, Liu X, et al. AMPKα modulation in cancer progression: multilayer integrative analysis of the whole transcriptome in Asian gastric cancer. Cancer research, 2012, 72(10): 2512-2521.
20. Treasure T, Fallowfield L, Lees B, et al. Pulmonary metastasectomy in colorectal cancer: the PulMiCC trial. Thorax, 2012, 67(2): 185-187.
21. Zhang L, Guo X, Zhang D, et al. Upregulated miR-132 in RESEARCH http://www.antpublisher.com/index.php/APT/index Lgr5+ gastric cancer stem cell-like cells contributes to cisplatin-resistance via SIRT1/CREB/ABCG2 signaling pathway. Molecular carcinogenesis, 2017, 56(9): 2022- 2034.
22. Trapasso S, Allegra E. Role of CD44 as a marker of cancer stem cells in head and neck cancer. Biologics: targets & therapy, 2012, 6: 379.
23. Sun X, Yang Z, Li S, et al. In vivo stabilization of polyethylene glycol (PEG)-modified recombinant methioninase (rMETase) activity by pyridoxal phosphate. 2004.
24. Tan Y, Sun X, Xu M, et al. Efficacy of recombinant methioninase in combination with cisplatin on human colon tumors in nude mice. Clinical Cancer Research, 1999, 5(8): 2157-2163.
25. Xin L, Cao J Q, Liu C, et al. Evaluation of rMETase-loaded stealth PLGA/liposomes modified with anti-CAGE scFV for treatment of gastric carcinoma. Journal of biomedical nanotechnology, 2015, 11(7): 1153-1161.
26. Aune T M, Spurlock III C F. Long non-coding RNAs in innate and adaptive immunity. Virus research, 2016, 212: 146-160.
27. Liu H, Wang G, Yang L, et al. Knockdown of long noncoding RNA UCA1 increases the tamoxifen sensitivity of breast cancer cells through inhibition of Wnt/β-catenin pathway. PloS one, 2016, 11(12).
28. Sun W, Yuan X, Tian Y, et al. Non-invasive approaches to monitor EGFR-TKI treatment in non-small-cell lung cancer. Journal of hematology & oncology, 2015, 8(1): 95.
29. Habib S L, Kasinath B S, Arya R R, et al. Novel mechanism of reducing tumourigenesis: upregulation of the DNA repair enzyme OGG1 by rapamycin-mediated AMPK activation and mTOR inhibition. European Journal of Cancer, 2010, 46(15): 2806-2820.
30. Ma Y, Zheng Y, Liu K, et al. Nanoparticles of poly (lactideco-glycolide)-da-tocopheryl polyethylene glycol 1000 succinate random copolymer for cancer treatment. Nanoscale research letters, 2010, 5(7): 1161-1169.
31. Miladi K, Ibraheem D, Iqbal M, et al. Particles from preformed polymers as carriers for drug delivery. EXCLI journal, 2014, 13: 28.
32. Xu Q, Hashimoto M, Dang T T, et al. Preparation of monodisperse biodegradable polymer microparticles using a microfluidic flow-focusing device for controlled drug delivery. Small, 2009, 5(13): 1575-1581.
33. Ayatollahi S, Hashemi M, Kazemi Oskuee R, et al. Synthesis of efficient gene delivery systems by grafting pegylated alkylcarboxylate chains to PAMAM dendrimers: evaluation of transfection efficiency and cytotoxicity in cancerous and mesenchymal stem cells. Journal of biomaterials applications, 2015, 30(5): 632-648.
34. Sheardown H, Van Beek M, Guo J. Hyaluronic acid-retaining polymers: U.S. Patent 7,674,781. 2010-3-9.