Open Access | Commentary
This work is licensed under a Creative Commons Attribution-ShareAlike 4.0 International License.
Machine learning paves the way forward for geropathology as- sessment
* Corresponding author: Warren Ladiges
Mailing address: Department of Comparative Medicine, School
of Medicine, University of Washington, Seattle, WA 98195, USA.
Email: wladiges@uw.edu
Received: 31 May 2024 / Accepted: 03 June 2024 / Published: 27 June 2024
DOI: 10.31491/APT.2024.06.142
Abstract
Geropathology investigates the pathological aspects of aging, analyzing age-related lesions in organs to track their progression and link to comorbidities. Utilizing geropathology grading scores, researchers evaluate drug efficacy in mammalian models, providing insights into the biological pathways of aging and disease development. Sheehan et al. introduced a novel approach using a weakly supervised machine learning algorithm to analyze age-related differences in mouse kidneys. This method, using chronological age as labels, effectively identifies age-associated histopathology. The model was validated against the Geropathology Research Network grading scheme, demonstrating high reliability and effectiveness in discerning drug treatment effects. The model’s reproducibility and ease of implementation, supported by tutorials and GitHub resources, enhance its utility. However, challenges include potential over-precision, the need for pathologist support, and strict whole slide image processing to prevent false detections. Future research could extend to additional systemic organs and various animal models, leveraging genetic similarities in lesion structures to improve grading precision and translational relevance to human aging and diseases.
Keywords
Geropathology, aging, machine learning, age-related lesions, drug efficacy
Geropathology, within the field of geroscience, focuses on
the pathological aspects of aging and its association with
diseases [1]. It involves assessing age-related lesions in
organs, grading their severity to assign quantitative values.
This detailed histopathological analysis helps track how
these lesions progress with age and their links to comorbidities. By using geropathology grading scores, researchers can evaluate the effectiveness of drugs in mammalian
animal models, providing crucial insights into the biological pathways of aging and the development of age-related
diseases. These grading scores serve as critical endpoints
in drug studies, allowing for precise measurement of how
treatments impact the severity and progression of ageassociated lesions.
Sheehan et al. recently introduced a novel approach, leveraging a supervised machine learning (ML) algorithm
to analyze age-related differences in mouse kidneys [2].
Unlike traditional ML methods, weakly supervised learning operates with loosely specified prediction targets,
blending aspects of both supervised and unsupervised
learning. Essentially, it discerns age-associated histopathology using sample-level chronological age for each
whole slide image (WSI), where higher chronological age
serves as the label. Analogous to teaching a student with
incomplete information, weakly supervised learning provides general hints or clues to the computer, prompting it
to discern patterns and make predictions based on these
cues. While not as precise as fully supervised learning,
which requires detailed labels for every data point, weakly
supervised learning empowers the computer to make educated guesses, providing invaluable information for tasks
where obtaining precise labels for all data are challenging
or time-consuming.
Sheehan et al. validated their model by correlating scores
generated by pathologists using the Geropathology Research Network aging grading scheme [3], demonstrating
high inter-reliability when compared to independent scoring by board-certified veterinary pathologists [2]. Furthermore, they showed that this classifier can discern not only
age-related differences, but also differences resulting from
drug treatment, such as a combination of rapamycin, acarbose, and phenylbutyrate, which was shown by our laboratory to be effective in enhancing resilience to aging in
C57Bl/6 and HET3 mice [4] (Figure 1). Importantly, the
model’s reproducibility allows for repeatable implementation in further studies, and the authors have provided
tutorials and step-by-step directions on how to implement the code provided on GitHub. Additionally, the classifier’s
throughput and portability make it feasible to run on individual computers.
Nevertheless, implementing this technique presents several significant challenges. As addressed by Sheehan et
al., while the evaluation of all pixels in an image ensures
unbiased scoring, it also risks over-precision of estimates.
Since the algorithm lacks the ability to differentiate lesions from non-lesions, its role is limited to identifying
differences between age groups. Thus, ongoing support
from trained pathologists is essential to discern subtle
details over time before full autonomy can be achieved.
Moreover, strict processing of WSIs is imperative to
maintain precision. Any mechanical variability in tissue
processing, slide preparation or imaging can significantly
impact the detection of false positives and negatives [5].
Standardizing these procedures is therefore crucial to
minimize variability. Lastly, this technique only detects
differences from normal tissue rather than specific abnormalities, unlike supervised ML methods. While valuable
for assessing age-related degeneration in organ systems, it
cannot replace supervised ML techniques for identifying
specific anomalies in tissues.
Further investigations could explore examining additional
systemic organs (i.e., brain, heart, lung, liver, pancreas,
spleen, skeletal muscle) within and beyond the mouse
model, extending to vertebrate models such as cats and
invertebrate models like the house cricket. Both hold
substantial translational significance in studying human
aging and age-related diseases and are of great interest for
our laboratory [6, 7]. Through leveraging the genetically
preserved similarities in lesion structure and shape across
species, machine learning techniques could be employed
to discern subtle differences between animal tissue types
and histopathological appearances. This approach would
not only enhance the precision of grading, but also shed
light on variations between animal models. Identifying
limitations and similarities in these models would allow
for more accurate interpretation of findings and better
understanding of translational relevance in histopathology
research for humans.
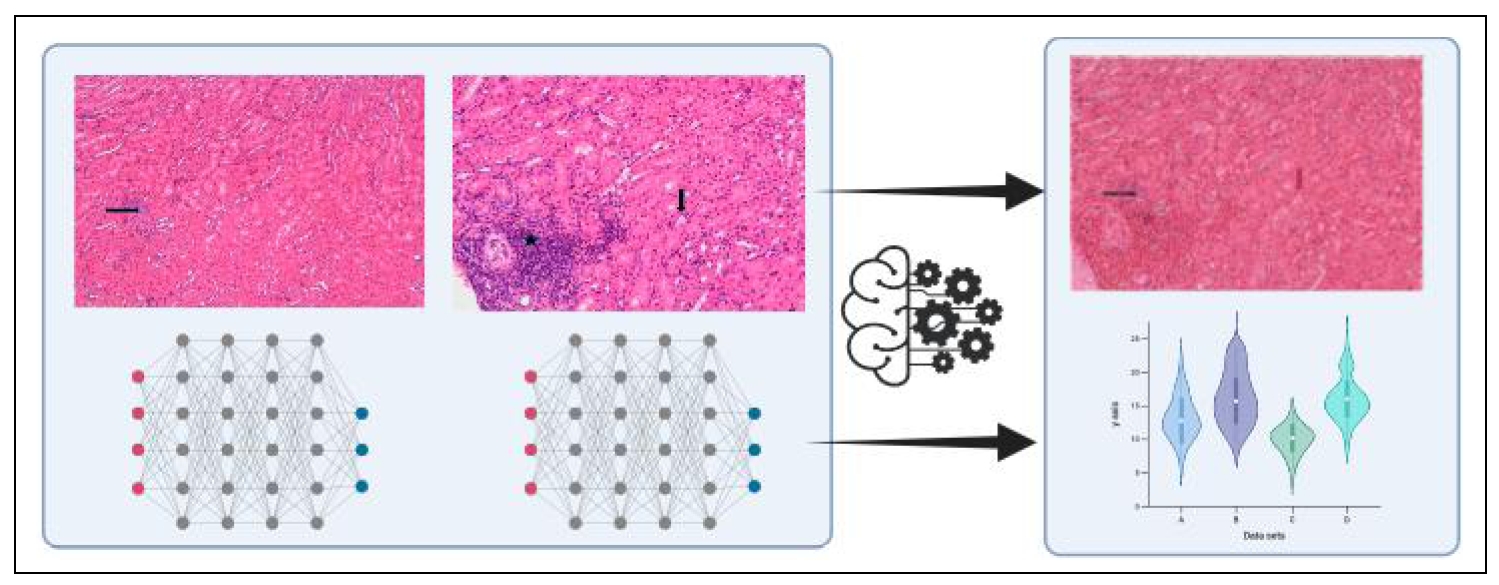
Figure 1. Schematic diagram illustrates the program utilizing deep neural network algorithms, which employ multiple layers of interconnected digital points to model complex patterns in data for image recognition. This program identifies and distinguishes between treatment and control cohorts for kidney analysis. The leftmost hematoxylin and eosin (H&E)-stained image demonstrates a kidney with grade 1 nephropathy, characterized by less than 10% of renal parenchyma affected by an age-related lesion, in this case focal tubular regeneration (black arrow), after drug treatment. The middle H&E image shows a kidney with grade 3 nephropathy characterized by moderate lesions affecting 30 to 70% parenchyma compared to grade 1, in this example demonstrated by tubular proteinuria (black arrow), as well as focal minimal lymphoid aggregates (grade 1, black star). Lesion categories and scores were assigned by two independent pathologists. The rightmost image shows how the system codes these differences by overlapping images and analyzing age-related variations, creating quantitative measurements that are displayed graphically. Magnification 20X.
Declarations
Financial support and sponsorship
None.
Conflict of interest statement
Warren Ladiges is a member of the editorial board of Aging Pathobiology and Therapeutics. The authors declare that they have no conflicts and were not involved in the journal’s review or decision regarding this manuscript.
References
1. Ladiges W, Ikeno Y, Niedernhofer L, McIndoe RA, Ciol MA, Ritchey J, et al. The geropathology research network: an interdisciplinary approach for integrating pathology into research on aging. J Gerontol A Biol Sci Med Sci, 2016, 71(4): 431-434. [Crossref]
2. Sheehan S, Mawe S, Chen M, Klug J, Ladiges W, Korstanje R, et al. A machine learning approach for quantifying age-related histological changes in the mouse kidney. Geroscience, 2024, 46(2): 2571-2581. [Crossref]
3. Snyder JM, Snider TA, Ciol MA, Wilkinson JE, Imai DM, Casey KM, et al. Validation of a geropathology grading system for aging mouse studies. Geroscience, 2019, 41(4): 455-465. [Crossref]
4. Jiang Z, Wang J, Imai D, Snider T, Klug J, Mangalindan R, et al. Short term treatment with a cocktail of rapamycin, acarbose and phenylbutyrate delays aging phenotypes in mice. Sci Rep, 2022, 12(1): 7300-7310. [Crossref]
5. La Perle KMD. Machine learning and veterinary pathology: be not afraid! Vet Pathol, 2019, 56(4): 506-507. [Crossref]
6. Wezeman J, Keely A, & Ladiges W. Resilience to aging drives personalized intervention strategies for Alzheimer’s disease. Aging Pathobiology and Therapeutics, 2023, 5(4): 151-153. [Crossref]
7. Liao GY, Rosenfeld M, Wezeman J, & Ladiges W. The house cricket is an unrecognized but potentially powerful model for aging intervention studies. Aging Pathobiology and Therapeutics, 2024, 6(1): 39-41. [Crossref]